Simon Roux
Research Scientist, Viral (Meta)Genomics
DOE Joint Genome Institute, Lawrence Berkeley National Laboratory
Biography
At the DOE Joint Genome Institute, I lead the Viral Genomics group where we explore viruses of microbes and their impacts on ecosystems using (mostly) fancy ‘omics tools. Our current projects include the study of viral diversity and virus:host interactions in soil and freshwater environments, along with the development of new bioinformatics tools and experimental protocols to probe and characterize uncultivated viruses. We also assist users of the JGI Metagenome Program with their analysis, including identification of viral sequences, functional annotation, taxonomic classification, etc.
The long-term goal of my research is to understand the ecological and evolutionary drivers of virus:host dynamics in natural microbial communities. This research involves a mix of experimental and computational approaches spanning from the molecular to the ecosystem scale, trying to address fundamental questions like “how do viruses spread and adapt across environments ?”, “how do viruses take over and reprogram microbial cells ?”, and “how do viral infections alter ecosystem processes ?”.
Interests
- Microbial/Viral Ecology
- (Viral) Metagenomics
- Virus Evolution
Education
-
PhD in Microbial Ecology, 2013
Université Blaise Pascal, Clermont-Ferrand II (now Université Clermont Auvergne)
-
MSc in Data Analysis and Modeling for Life Sciences, 2010
Université Blaise Pascal, Clermont-Ferrand II (now Université Clermont Auvergne)
-
BSc in Microbiology, 2008
Université Blaise Pascal, Clermont-Ferrand II (now Université Clermont Auvergne)
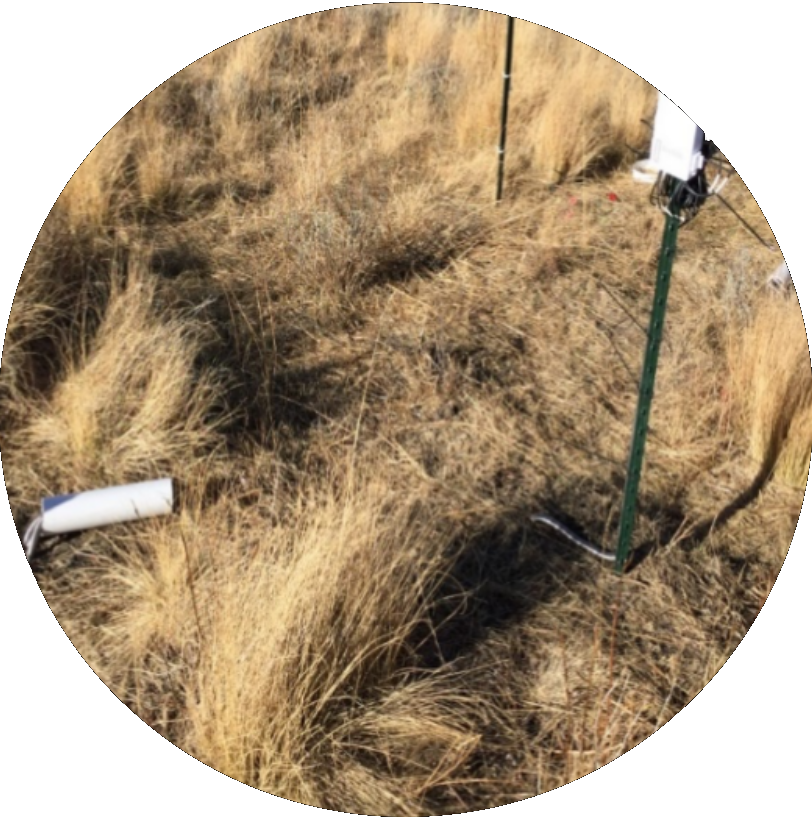 -
Virus-driven alterations of microbial metabolism in soil
-
Virus-driven alterations of microbial metabolism in soil  -
Virus:host dynamics in Yellowstone National Park biofilms
-
Virus:host dynamics in Yellowstone National Park biofilms  -
Defining the healthy human virome
-
Defining the healthy human virome  -
Establishing the foundations of a high-throughput phage foundry
-
Establishing the foundations of a high-throughput phage foundry  -
IMG/VR - Large-scale exploration of uncultivated viral diversity
-
IMG/VR - Large-scale exploration of uncultivated viral diversity